Figure 4: Metal ion interactions.
From Radical SAM enzyme QueE defines a new minimal core fold and metal-dependent mechanism
- Daniel P Dowling1, 2,
- Nathan A Bruender3,
- Anthony P Young3,
- Reid M McCarty3,
- Vahe Bandarian3,
- Catherine L Drennan1, 2, 4,
- Journal name:
- Nature Chemical Biology
- Volume:
- 10,
- Pages:
- 106–112
- Year published:
- DOI:
- doi:10.1038/nchembio.1426
- Received
- Accepted
- Published online
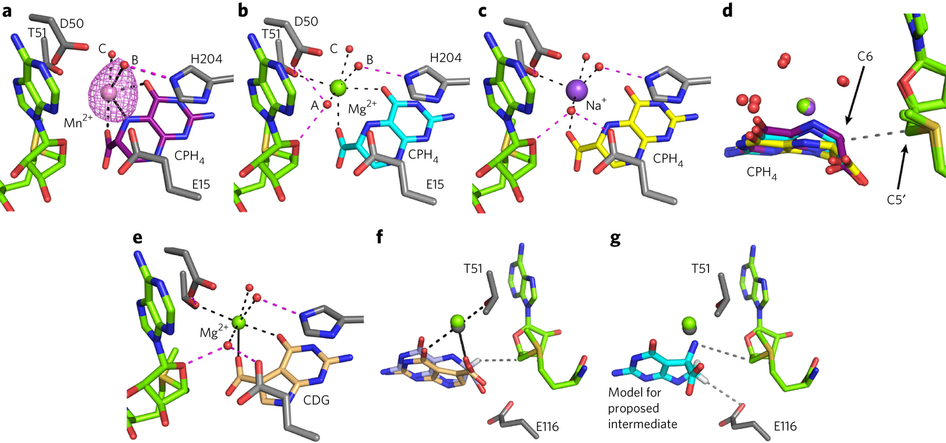
(a–c) QueE structures with CPH4 and different metals, colored as follows: Mn2+/CPH4, pink/purple (a); Mg2+/CPH4, green/cyan (b); and Na+/CPH4, purple/yellow (c). Hydrogen bonding interactions with metal-bound water molecules are displayed as magenta dashed lines, coordination distances are shown as solid black lines, and metal-ligand distances between 2.5 and 3.2 Å are represented by dotted black lines. The distances are presented in Supplementary Table 4. Anomalous difference electron density for Mn2+ is contoured at 3σ (pink mesh). AdoMet carbons, green; protein carbons, gray; water, red spheres. (d) An overlay of all three CPH4-bound structures colored as in a–c. The distance between C6 of the substrate and the C5′ of AdoMet is represented as a dashed gray line. (e) QueE cocrystallized with Mg2+/CDG in green/tan; the lines and other colors are as in a. (f) A superposition of substrate- and product-bound QueE colored as in a, with Mg2+/CPH4 and Mg2+/CDG in green/light blue and gray/tan, respectively. The distance (3.4 Å) between the AdoMet C5′ and the CPH4 C6 H atom is displayed as a dashed gray line. (g) The proposed exocyclic nitrogen-containing intermediate is modeled into the QueE active site with cyan carbons based on the binding mode of CPH4 to preserve reasonable distances to the metal ion–binding site. This orientation positions the proposed nitrogen radical within reasonable distance (∼4 Å) to the C5′ of AdoMet, displayed as a dashed gray line, for reabstraction of an H atom to regenerate the AdoMet cofactor. The distance (∼3.5 Å) between the C7 proton to be lost in the reaction and the nearest protein residue, E116, is also represented as a dashed gray line.
Additional data
Entities in this article
-
7-carboxy-7-deazaguanine synthase QueE
Bmul_3115
Burkholderia multivorans (strain ATCC 17616 / 249)
View: -
7-carboxy-7-deazaguanine synthase QueE
queE
Bacillus subtilis (strain 168)
View: -
Ferredoxin--NADP reductase
fpr
Escherichia coli (strain K12)
View: -
Pyruvate formate-lyase 1-activating enzyme
pflA
Escherichia coli (strain K12)
View: -
Formate acetyltransferase 1
pflB
Escherichia coli (strain K12)
View: -
Biotin synthetase HmdB
bioB
Methanococcus maripaludis (strain S2 / LL)
View: -
GTP cyclohydrolase FolE2
BMULJ_03697
Burkholderia multivorans (strain ATCC 17616 / 249)
View: -
6-pyruvoyl tetrahydrobiopterin synthase QueD
Bmul_3114
Burkholderia multivorans (strain ATCC 17616 / 249)
View: -
7-cyano-7-deazaguanine synthase
exsB
Burkholderia multivorans (strain ATCC 17616 / 249)
View: -
cysteine
View: -
6-carboxy-5,6,7,8-tetrahydropterin
View: -
7-carboxy-7-deazaguanine
View: -
ammonia
View: -
purine
View: -
guanosine 5'-triphosphate
View: -
7-deazapurine
View: -
dithionite
View: -
6-carboxypterin
View: -
methionine
View: -
ribose
View: -
adenine
View: -
pterin
View: -
tetrahydropterin
View: -
manganese(II) sulfate
View: -
magnesium(II) chloride
View: -
water
View: -
threonine
View: -
5′-deoxyadenosine
View: -
tetrahydropyrazine
View: -
pyrrole
View: -
kanamycin
View: -
ampicillin
View: -
arabinose
View: -
iron(III) chloride
View: -
isopropyl β-D-1-thiogalactopyranoside
View: -
hydrogen
View: -
potassium chloride
View: -
imidazole
View: -
phenylmethylsulfonyl fluoride
View: -
nickel sulfate
View: -
piperazine-N,N'-bis(2-ethanesulfonic acid)
View: -
dithiothreitol
View: -
sodium sulfide
View: -
magnesium(II) sulfate
View: -
trichloroacetic acid
View: -
trifluoroacetic acid
View: -
acetonitrile
View: -
NADPH
View: -
sodium dipotassium phosphate
View: -
sodium acetate
View: -
glycerol
View: -
acetate
View: -
4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
View: -
glycine
View: -
proline
View: -
preQ0
View: -
adenosine 5'-triphosphate
View: -
toyocamycin
View:
Affiliations
-
Howard Hughes Medical Institute, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
- Daniel P Dowling &
- Catherine L Drennan
-
Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
- Daniel P Dowling &
- Catherine L Drennan
-
Department of Chemistry and Biochemistry, University of Arizona, Tucson, Arizona, USA.
- Nathan A Bruender,
- Anthony P Young,
- Reid M McCarty &
- Vahe Bandarian
-
Department of Biology, Massachusetts Institute of Technology, Cambridge, Massachusetts, USA.
- Catherine L Drennan
Contributions
D.P.D. and C.L.D. designed and performed the crystallography experiments. N.A.B., R.M.M., A.P.Y. and V.B. designed and carried out the biochemical experiments. D.P.D., N.A.B., V.B. and C.L.D. contributed to the writing of the manuscript.
Competing financial interests
The authors declare no competing financial interests.
Author details
Daniel P Dowling
Search for this author in:
Nathan A Bruender
Search for this author in:
Anthony P Young
Search for this author in:
Reid M McCarty
Search for this author in:
Vahe Bandarian
Search for this author in:
Catherine L Drennan
Search for this author in:

